- 🌱 Biotic interactions play a key role in shaping species distributions
- 📊 Incorporating biotic variables into species distribution models (SDMs) is therefore essential
- ⚠️ However, biotic data are often incomplete or inconsistently available across species and locations
🎯Our goal: leverage incomplete biotic information during both training and inference
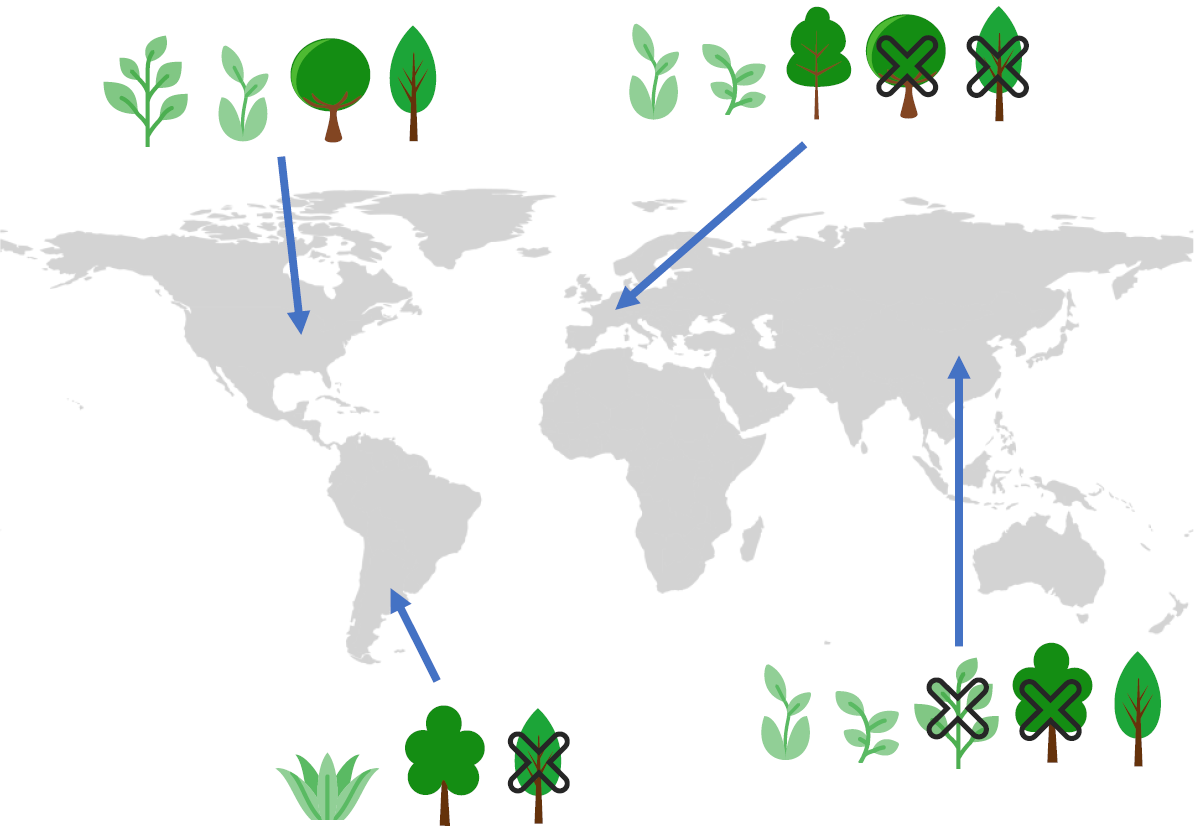
🚀 We propose CISO, a multi-species deep learning approach for SDMs that can be conditioned on any available species presence or absence
Abstract
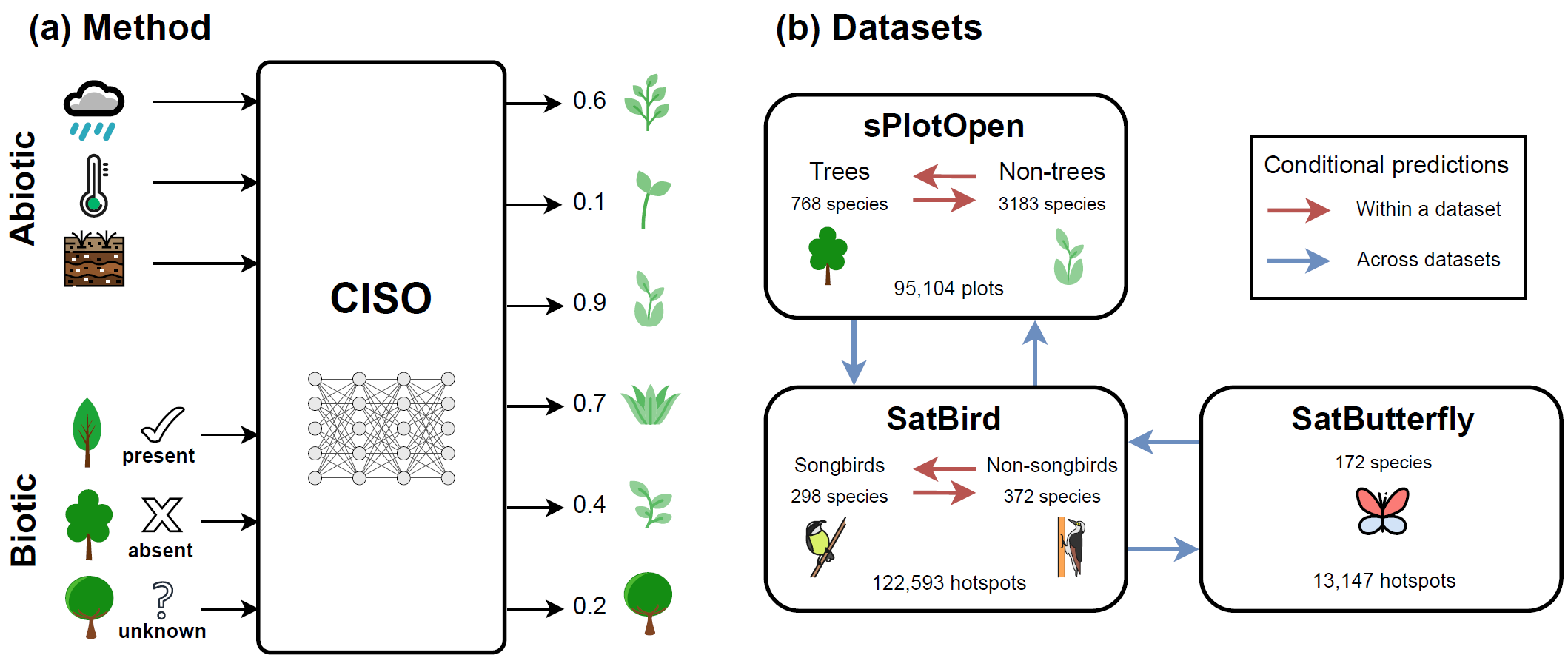
Species distribution models (SDMs) are widely used to predict species' geographic distributions, serving as critical tools for ecological research and conservation planning. Typically, SDMs relate species occurrences to environmental variables representing abiotic factors, such as temperature, precipitation, and soil properties. However, species distributions are also strongly influenced by biotic interactions with other species, which are often overlooked in traditional models. While some methods, such as joint species distribution models (JSDMs), partially address this limitation by incorporating biotic interactions, they often assume symmetrical pairwise relationships between species and require consistent co-occurrence data. In practice, species observations are often sparse, and the availability of information about the presence or absence of other species varies significantly across locations. To address these challenges, we propose CISO, a deep learning-based method for species distribution modeling Conditioned on Incomplete Species Observations. CISO enables predictions to be conditioned on a flexible number of species observations alongside environmental variables, accommodating the variability and incompleteness of available biotic data. We demonstrate our approach using three datasets representing different species groups: sPlotOpen for plants, SatBird for birds, and a new dataset, SatButterfly, for butterflies. Our results show that including partial biotic information improves predictive performance on spatially separate test sets. When conditioned on a subset of species within the same dataset, CISO outperforms alternative methods in predicting the distribution of the remaining species, for plants and birds. Furthermore, we show that combining and conditioning on observations from multiple datasets can improve the prediction of species occurrences in scenarios with sufficient co-occurrences between datasets to train CISO effectively. Our results show that CISO is a promising ecological tool, capable of incorporating incomplete biotic information and identifying potential interactions between species from disparate taxa.
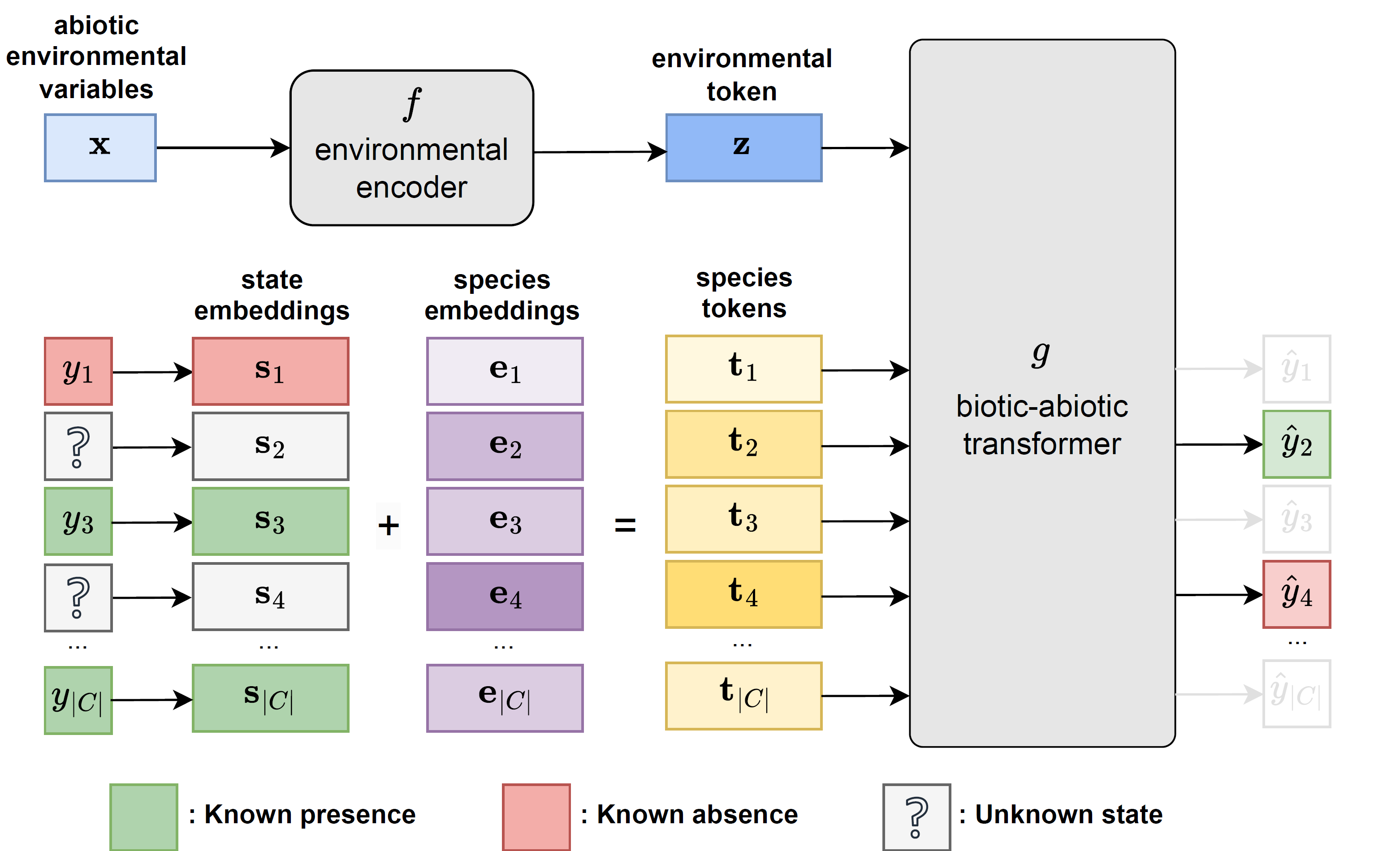
CISO architecture overview
- 🧩 Environmental variables and available species observations are encoded as tokens and processed by a transformer, enabling non-linear combination of abiotic and biotic information
- 🎭 During training, species observations are randomly masked to simulate incomplete observation scenarios (masked modeling). The model learns to predict the presence or absence of the remaining species
- 🔍 At inference time, CISO can be conditioned on any subset of available species observations to predict the occurrence of target species
📄 Check out the paper and 💻 code for more details!
Citation
@article{abdelwahed2026ciso,
title={CISO: Species Distribution Modeling Conditioned on Incomplete Species Observations},
author={Abdelwahed, Hager Radi and Teng, M{\'e}lisande and Zbinden, Robin and Pollock, Laura and Larochelle, Hugo and Tuia, Devis and Rolnick, David},
journal={Methods in Ecology and Evolution},
year={2026}
}





